Recent Studies
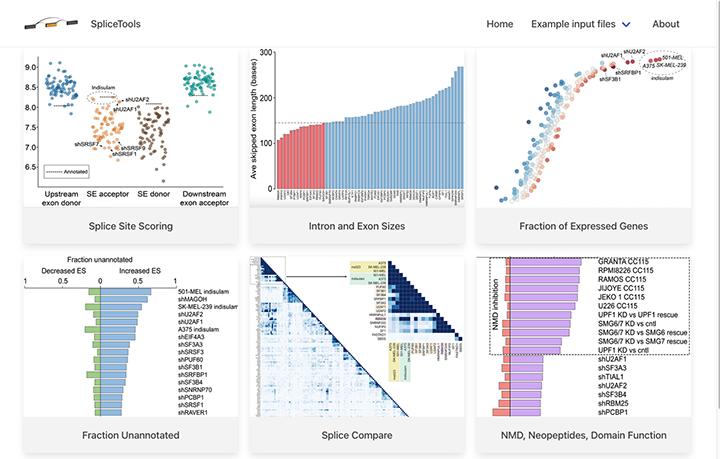
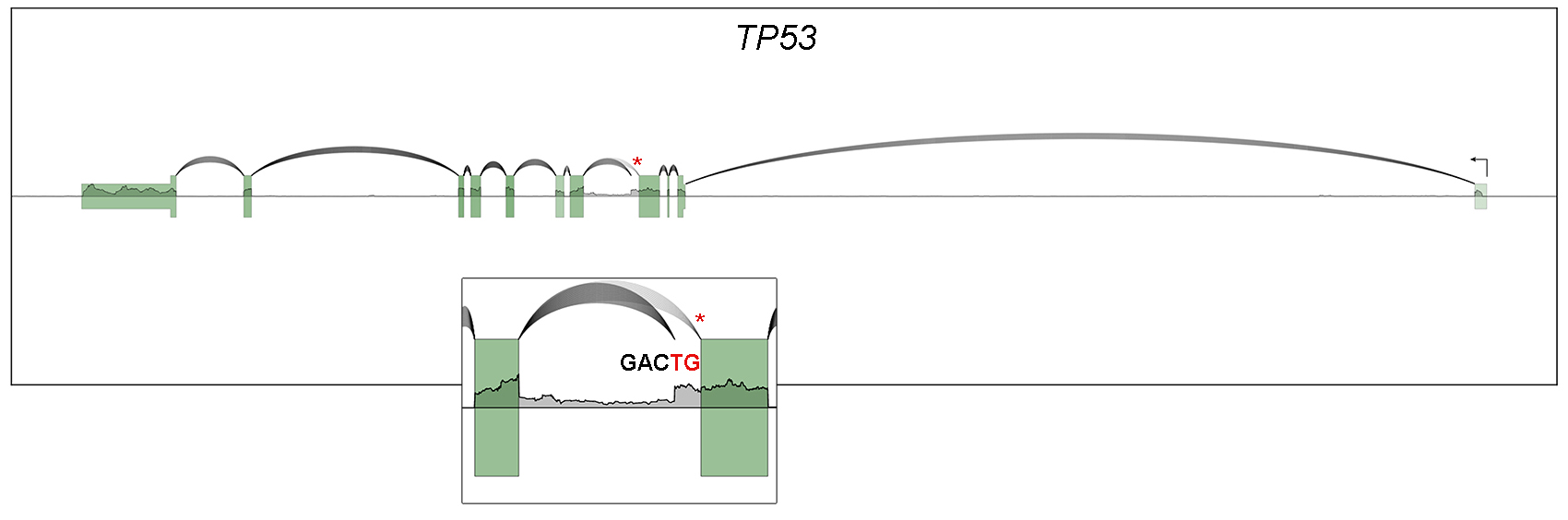
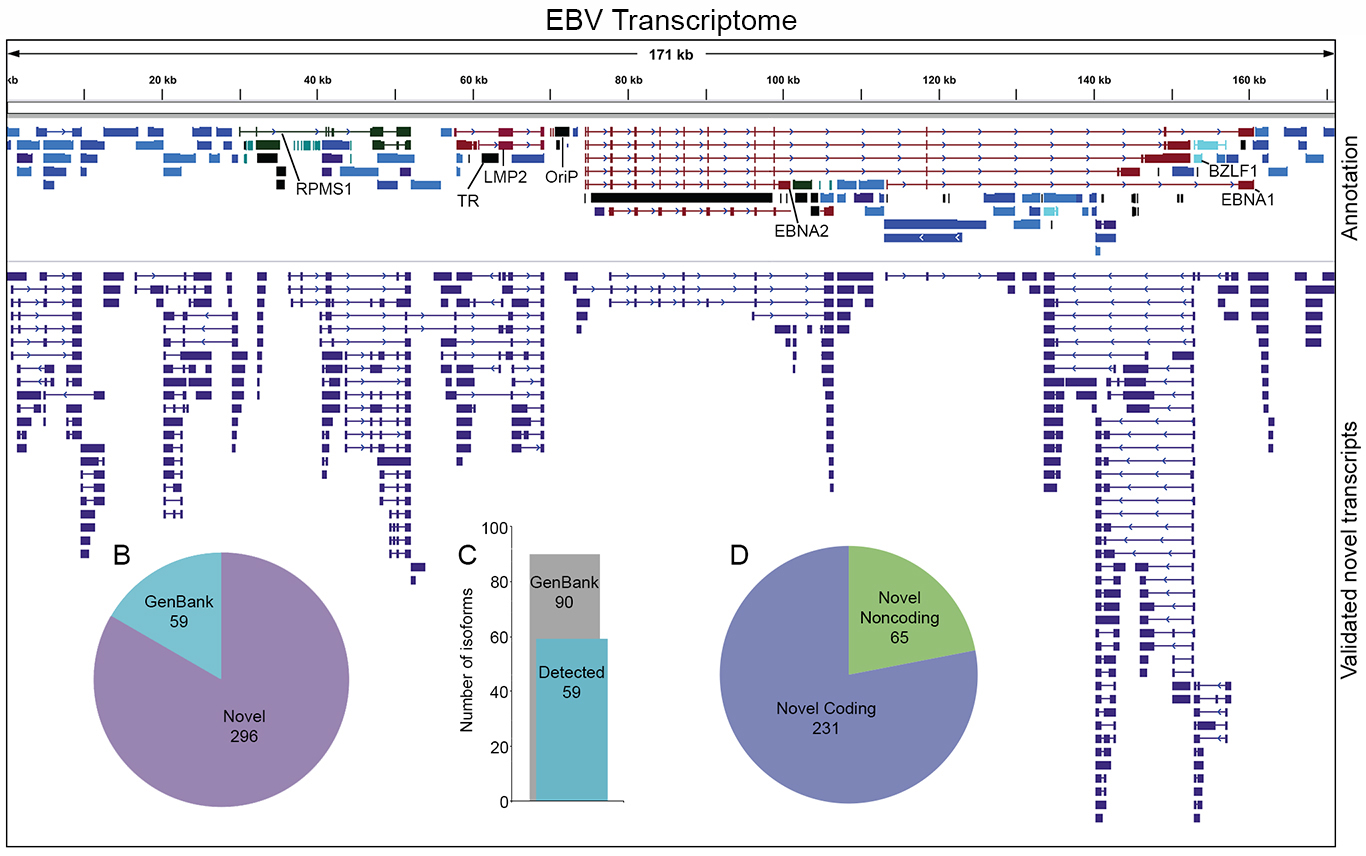
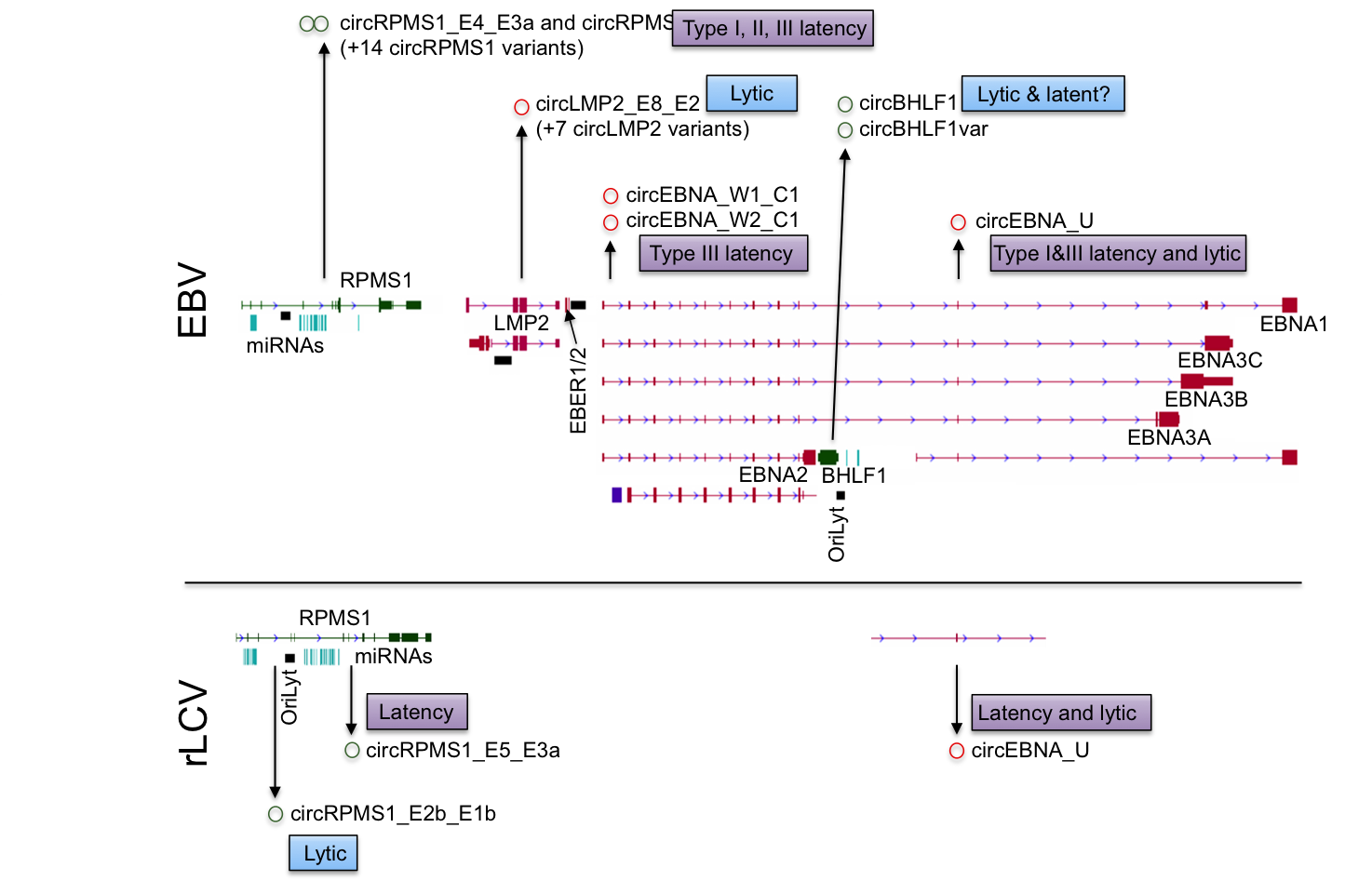
Recent studies from the Flemington lab include the development of a suite of RNA splicing analysis tools (Flemington et al, Nucleic Acids Research 2023), the development of data visualization and analysis software (e.g. Ungerleider and Flemington, BMC Bioinformatics 2019), the development of software to globally resolve complex transcript structures in high gene density genomes (O’Grady et al, Nucleic Acids Res, 2016), the discovery of new transcripts and transcript isoforms in EBV (O’Grady, Nucleic Acids Res, 2016) and murine herpesvirus 68 (MHV68, O’Grady et al, Cell Reports 2019), and the discovery of circular RNAs encoded by EBV (Ungerleider et al, Plos Pathogens 2018) and rhesus lymphocryptovirus (rLCV), Kaposi’s Sarcoma Herpesvirus (KSHV), and murine herpesvirus 68 (MHV68, Ungerleider et al, J Virol 2019 and Ungerleider et al, mBio 2019).
Current Focus
The current focus of the Flemington lab involves investigating changes in host cell environment following EBV infection/reactivation. We are using a multi-omics approach to study the multi-functional roles of specific EBV early lytic genes on the cell transcriptome, cell chromosome reorganization, and NMD (nonsense mediated decay) and studying EBV microRNAs and their role in Burkitt’s Lymphoma progression. We are also employing Machine Learning to predict motifs related to de novo promoter cell transcription and the control of cellular gene expression.